Basic Tuberculosis Research

Project Lead(s)

Professor

Senior Research Fellow
Project Background
Before the 19th century, very little was known about the causative pathogen and disease mechanisms. Now we know that tuberculosis in mammals is mainly caused by at least 9 genetically related mycobacterial species comprising Mycobacterium tuberculosis sensu stricto (Mtbss), M. africanum (Maf), M. bovis, M. mungi, M. microti, M. caprae, M. pinnipedii, M. suricattae, and M. orygis together referred to as the M. tuberculosis complex (MTBC). Previously thought to be a monomorphic bacterium, current application of next generation sequencing tools has revealed substantial genomic differences within members of the complex. In our lab, our basic TB research aims to understand the biology and transmission of the causative pathogen as well as gain insight into understanding host-pathogen interactions using various approaches comprising basic molecular epidemiology, genomic epidemiology, comparative genomics, phenotypic and genotypic analysis, and immunology.
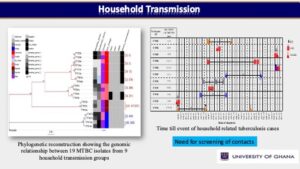
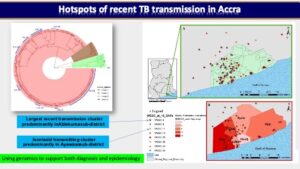
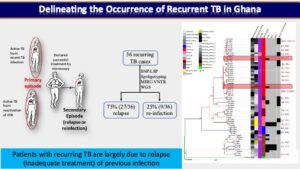
In one of our prospective cohort-based research, we used a population-based molecular epidemiology study to determine the transmission pattern and dynamics of Mycobacterium tuberculosis complex (MTBC) strains from two areas in Ghana (East Mamprusi and Accra Metropolitan Area) and identify associated risk factors to complement conventional control program efforts. We used SaTScan and ArcMap analyses to identify and map significantly clustered MTBC lineages/sub-lineages. Isolates were strain typed using standard mycobacterial interspersed repetitive-unit variable-number tandem repeat (MIRU-VNTR) typing and then ultimately whole genome sequencing (WGS) while performing phylogenetic analysis to assess strain relatedness. Among 2,551 isolates genotyped for spatio-temporal analysis, 2,019 (79.1%), 516 (20.2%) and 16 (0.6%) were identified as Mtbss, Maf and animal strains respectively. Maf lineages were found to persist at approximately 20% over an 8-year period. Whereas the Cameroon sub-lineage was associated with Southern Ghana, the Beijing, Ghana and animal genotypes were significantly (p<0.05) associated with Northern Ghana. We estimated recent transmission rate of 24.7% using WGS and confirmed the existence of reduced recent transmission of Maf compared to Mtbss L4. Observed reduced recent transmission of Maf suggests other factor(s) (host/environmental) may be responsible for its continuous presence in West Africa. WGS combined with epidemiological information confirmed a wide spread of a Cameroon sub-lineage clone mainly from the Ablekuma sub-district. More importantly, we identified a recent transmission cluster associated with isoniazid resistance belonging to the Ghana sub-lineage of L4. Risk factor analysis using logistic regression modeling, identified younger individuals (age <30 years) and male gender as significant risk factors for recent TB transmission. Majority (94.4%, 34/36) of individuals with recurring TB episodes were males and 58.6% (21/36) had TB recurrence within 12 months post treatment. Recurrent TB was attributed to inadequate treatment (relapse of same strain) in 75.0% (27/36) of participants with 25.0% (9/36) attributed to re-infection. Epidemiologically linked TB cases were likely the results of recent TB transmission within the house or from neighboring recent transmission events. We show that it is possible to monitor recurring TB cases and follow-up household related transmission in a resource-limited setting.
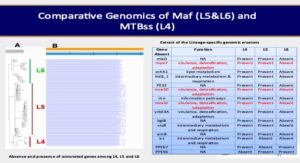
To understand further and add to knowledge the biology of the human adapted MTBC (hMTBC),ie, Mtbss and Maf, we conducted a study to determine the genomic diversity among the dominant MTBC lineages (L) (L4, L5 and L6) in Ghana and the functional implications of the observed genomic variability. We observed that L6 was found to have accumulated significantly more fixed single nucleotide polymorphisms (SNPs) (median = 1037; p < 0.05) compared to L4 (median = 822) and L5 (median = 928) in addition to significantly higher pairwise nucleotide diversity (π). The whole genome phylogeny of Maf in Ghana upon clustering showed 5 and 3 clusters of L5 and L6 respectively. Average nucleotide identity (ANI) between any two lineages (L4/L5, L4/L6 and L5/L6) was always greater than 99%. The π of the functional genes was all significantly higher among the L6 compared to L4 and L5 with prominence among T cell epitopes (0.000061, 0.000063 and 0.000149 for L4, L5 and L6). Apart from T cell epitopes and regulatory proteins, all the gene families were under purifying selection thus the ration of change in nonsynonymous to change in synonymous mutations less than one (dN/dS <1.0) irrespective of the lineage. T cell epitopes of L4 and L5 were under purifying selection; thus the ratio of change in nonsynonymous to change in synonymous mutations (dN/dS) less than one (L4: median dN/dS = 0.57 and median dN/dS = 0.64 for fL4 and L5 respectively) compared to positive selection in L6 (median dN/dS = 1.53). Most T cell epitopes were conserved with only 10.4% (128/1226) mutated with lineage-specific variations. Genomic erosion was predicted to be more pronounced in L6 (3992 genes present) compared to L5 (4035 genes present) with predicted lineage-specific impaired expression of proteins such ESAT-6, mpt64 and a number of mce3 proteins. This study found the Ghana genotype of L4 to be associated with DR-TB in Ghana (p < 0.05) compared to the other dominant genotypes. INH resistant Mtbss and Maf were more likely kat G S315T and inhApro -15C/T mutants respectively. This study identified potential genotype-specific epistatic interaction DR-conferring and compensatory mutations. Finally, our conclusion is that, the hMTBC albeit closely related are genomically distinct. The West Africa restricted L5 and L6 together classified as Maf are genomically two different biovars of the MTBC, so different from each other as they are from L4, suggesting different biovars of Maf with potential sub-lineages. The observed genomic difference among the lineages has potential implications for virulence and applicability of TB control tools.
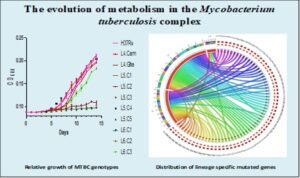
Genomic diversity among the causative agent of tuberculosis has not been considered in development of control tools. Most existing control tools were developed with knowledge of M. tuberculosis (Mtbss) to the neglect of M. africanum (Maf) (restricted to West Africa). Comparative genomics analysis of M. tuberculosis complex (MTBC) isolates (Lineages 4 to 6 and M. bovis) from Ghana (harbouring both Mtbss and Maf) has shown that the lineages of the MTBC are genomically distinct with lineage-specific genomic erosions, mutations and conservation of human T cell antigens and epitopes. The observed lineage-specific genomic events could possibly influence fitness, transmission potential, emergence of drug resistance and response to control tools. As part of efforts to explore the phenotypic implications of the observed genomic diversity of the MTBC in Ghana, this study intends to compare growth rate, gene expression and metabolome profiles of MTBC lineages under variable nutritional conditions. The work is ongoing but preliminary results show significant between lineage as well as within lineage phenotypic diversity among the MTBC with respect to their ability to utilize different sources of carbon and nitrogen.
Key Findings


Key Publications
- Asante-Poku, A., Morgan, P., Osei-Wusu, S., Aboagye, S. Y., Asare, P., Otchere, I. D., . . . Yeboah-Manu, D. (2022). Genetic Analysis of TB Susceptibility Variants in Ghana Reveals Candidate Protective Loci in SORBS2 and SCL11A1 Genes. Frontiers in Genetics, 12. doi:10.3389/fgene.2021.729737
- Osei-Wusu, S., Morgan, P., Asare, P., Adams, G., Musah, A. B., Siam, I. M., . . . Yeboah-Manu, D. (2021). Bacterial Load Comparison of the Three Main Lineages of Mycobacterium tuberculosis Complex in West Africa. Frontiers in Microbiology, 12(3176). doi:10.3389/fmicb.2021.719531
- Osei-Wusu, S., Otchere, I. D., Morgan, P., Musah, A. B., Siam, I. M., Asandem, D., . . . Yeboah-Manu, D. (2021). Genotypic and phenotypic diversity of Mycobacterium tuberculosis complex genotypes prevalent in West Africa. PloS One, 16(8), e0255433. doi:10.1371/journal.pone.0255433
- Asare, P., Osei-Wusu, S., Baddoo, N. A., Bedeley, E., Otchere, I. D., Brites, D., . . . Yeboah-Manu, D. (2021). Genomic epidemiological analysis identifies high relapse among individuals with recurring tuberculosis and provides evidence of recent household-related transmission of tuberculosis in Ghana. International Journal of Infectious Diseases, 106, 13-22. doi:10.1016/j.ijid.2021.02.110
- Asare, P., Otchere, I. D., Bedeley, E., Brites, D., Loiseau, C., Baddoo, N. A., . . . Yeboah-Manu, D. (2020). Whole Genome Sequencing and Spatial Analysis Identifies Recent Tuberculosis Transmission Hotspots in Ghana. Frontiers in Medicine, 7(161). doi:10.3389/fmed.2020.00161
- Otchere, I. D., Coscollá, M., Sánchez-Busó, L., Asante-Poku, A., Brites, D., Loiseau, C., . . . Yeboah-Manu, D. (2018). Comparative genomics of Mycobacterium africanum Lineage 5 and Lineage 6 from Ghana suggests distinct ecological niches. Scientific Reports, 8(1), 11269. doi:10.1038/s41598-018-29620-2
- Asare, P., Asante-Poku, A., Prah, D. A., Borrell, S., Osei-Wusu, S., Otchere, I. D., . . . Yeboah-Manu, D. (2018). Reduced transmission of Mycobacterium africanum compared to Mycobacterium tuberculosis in urban West Africa. International Journal of Infectious Diseases, 73, 30–42. doi:10.1016/j.ijid.2018.05.014
- Osei-Wusu, S., Amo Omari, M., Asante-Poku, A., Darko Otchere, I., Asare, P., Forson, A., . . . Yeboah-Manu, D. (2018). Second-line anti-tuberculosis drug resistance testing in Ghana identifies the first extensively drug-resistant tuberculosis case. Infection and Drug Resistance, Volume 11, 239-246. doi:10.2147/IDR.S152720
- Yeboah-Manu, D., de Jong, B. C., & Gehre, F. (2017). The Biology and Epidemiology of Mycobacterium africanum. In Adv Exp Med Biol (Vol. 1019, pp. 117-133).
- Otchere, I. D., Harris, S. R., Busso, S. L., Asante-Poku, A., Osei-Wusu, S., Koram, K., . . . Yeboah-Manu, D. (2016). The First population structure and comparative genomics analysis of Mycobacterium africanum strains from Ghana reveals higher diversity of Lineage 5. Paper presented at the Int J Mycobacteriol. https://www.ncbi.nlm.nih.gov/pubmed/28043631
- Stucki, D., Brites, D., Jeljeli, L., Coscolla, M., Liu, Q., Trauner, A., . . . Gagneux, S. (2016). Mycobacterium tuberculosis lineage 4 comprises globally distributed and geographically restricted sublineages. Nature Genetics, 48(12), 1535-1543. doi:10.1038/ng.3704
- Yeboah-Manu, D., Asare, P., Asante-Poku, A., Otchere, I. D., Osei-Wusu, S., Danso, E., . . . Gagneux, S. (2016). Spatio-Temporal Distribution of Mycobacterium tuberculosis Complex Strains in Ghana. PloS One, 11(8), e0161892. doi:10.1371/journal.pone.0161892
- Asante-Poku, A., Yeboah-Manu, D., Otchere, I. D., Aboagye, S. Y., Stucki, D., Hattendorf, J., . . . Gagneux, S. (2015). Mycobacterium africanum is associated with patient ethnicity in Ghana. PLoS Neglected Tropical Diseases, 9(1), e3370. doi:10.1371/journal.pntd.0003370
Team
Internal Collaborator(s)
Prof. Kwadwo Ansah Koram (MD, PhD)
External Collaborator(s)
Dr. George Kyei, UGMC
Dr. Gloria Ansa, University of Ghana Hospital
Dr. Audrey Forson, Consultant Pulmonologist, Chest Clinic, KBTH
Dr. Abraham Adjei, Chest Clinic, KBTH
Dr. Iddrisu Yahaya, Chest Clinic, KBTH
Dr. Yayra Klinogo, Chest Clinic, KBTH
Dr. Akosua Baddoo, Chest Clinic/NACP, KBTH
Dr. Jane Afriyie-Mensah, MD, Head Chest Dept
Dr. Yacoba Atiase, MD, Head Diabetic Clinic
Mr. Michael Amo Omari, Chest Clinic, KBTH
Mrs. Nelly Arthur, Chest Clinic, KBTH
Dr. Charlotte Alberta Cato, Mamprobi Polyclinic
Dr. Clement Laryea, 37-Millatary Hospital
Dr. Farida Abdulai, Regional TB Coordinator, GHS
Dr. Frank Bonsu, National TB control Programme (NTP), Ghana
Dr. Yaw Adusi-Poku, National TB control Programme (NTP), Ghana
Dr. Rita Patricia Frimpong, National TB control Programme (NTP), Ghana
Prof Martin Antonio, MRC Gambia
Dr Toyin Togun, London School of Hygiene and Tropical Medicine
Prof Sebastien Gagneux, Swiss Tropical and Public Health Institute
Prof Jakob Zingstag, Swiss Tropical and Public Health Institute
Prof Douglas Young, NIMR, MRC, UK
Dr Simon Harris, Wellcome Sanger Institute, UK
Prof. James Wood, Department of Veterinary Medicine, University of Cambridge, United Kingdom
Prof Julian Parkhill, Department of Veterinary Medicine, University of Cambridge, United Kingdom
Thomas Junghanss, Heidelberg, Germany, Africanum Consortium
Jacques Fellay, Ecole Polytechnique Fédérale de Lausanne (EPFL), Switzerland
Prof. Luiz de Carvalho, UK
Funder(s)
Ghana Health Service
Wellcome Trust
Wellcome Trust Sanger Institute
Swiss Tropical and Public Health Institute
Leverhulme-Royal Society Africa Award
CAN

